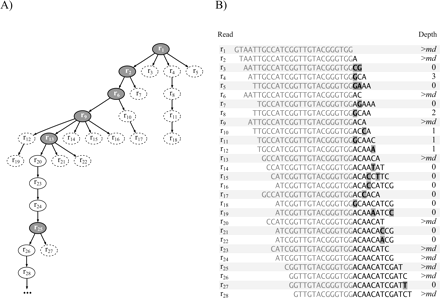
Removing short dead-end paths. (A) Possible path elongations from the right end of the read r1 are represented by a tree. Nodes that are removed are dashed. Each path leaving a branching node (shown in gray) is tested for the minimum depth it can initiate. If the required depth of md cannot be reached, then the nodes forming the dead-end path are removed. (B) Multiple sequence alignment of the reads belonging to the possible right end elongation of the read r1 is shown. The residues that do not agree with the consensus sequence are shaded. On the right side is indicated the depth value that can be reached by continuing the elongation from the corresponding read. The reads containing one or more mismatched residues have a low or a null depth value, indicating that no exact overlap exists for their right end in the entire reads data set. These reads are likely to contain sequencing errors.











