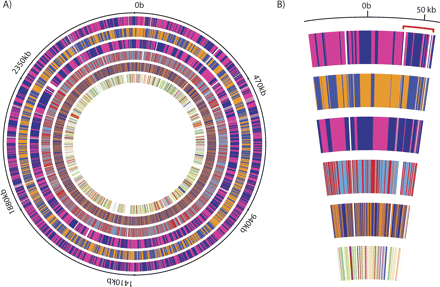
Mapping of the contigs on the reference Staphylococcus aureus MW2 genome. (A) From external to internal, the circles correspond to the contigs produced by (1) Edena strict, (2) Velvet, (3) Edena nonstrict, (4) SSAKE, and (5) SHARCGS. The contigs are colored by alternating two different colors, which allows distinguishing contig boundaries. The last inner circle shows the coding sequences. The gaps in the Edena nonstrict assembly correspond to large misassembled contigs that did not properly map the reference genome. (B) The magnification of the region around the origin of replication provides a better view to compare the contigs length and layout between the different assembly methods. It can be seen that the contigs assembled by Edena and Velvet are long enough to reveal entire genes. More importantly, significant overlaps exist between the contigs assembled by the two programs, which also means that even larger contigs could be assembled by merging both approaches. The position of the SSCmec cassette of type IV.1 (Chongtrakool et al. 2006) is indicated by the red line.











