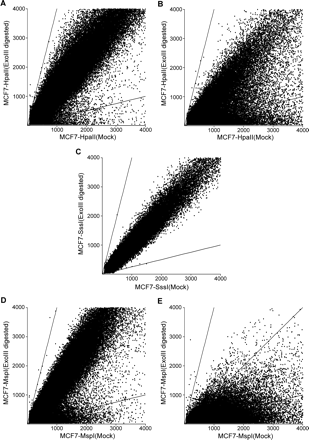
Scatter plot of TspRI–ExoIII–array CGH data. (A) MCF7 genomic DNA digested with TspRI–HpaII–ExoIII; hybridization signals of the data points not containing HpaII sites flanked by TspRI sites are close to a ratio of one between ExoIII-digested and control mock-digested DNA, as expected since these TspRI fragments should not contain ExoIII-sensitive ends. The two lines correspond to ratios of four and 0.25, respectively, between the Cy3 and Cy5 signals. Some of the HpaII-sensitive background is probably due to sequence polymorphism creating new HpaII sites, as revealed by SssI and MspI experiments. (B) A large fraction of hybridization signal in the 53,728 TspRI fraction containing HpaII sites is lower after digestion with HpaII and ExoIII. (C) After SssI methylation of CpG sites in vitro, the 53,728 TspRI fragments containing HpaII sites became resistant to HpaII–ExoIII digestion with a hybridization ratio of around one in contrast to the result in B. (D,E) MCF7 genomic DNA digested with CpG-methylation–insensitive restriction enzyme MspI results in depletion of signals with a ratio of one in the fraction containing HpaII sites and a lowering of signals, whereas the fragments without HpaII sites are not affected significantly.











