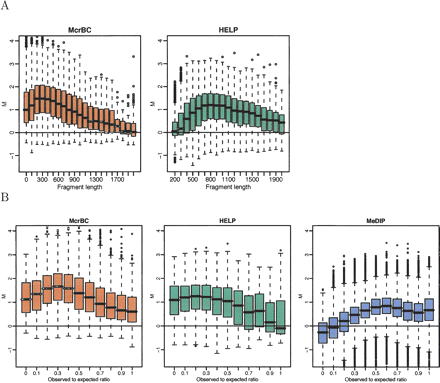
DNA fragment-length–related biases. (A) M-values for the HCT116 sample are stratified by the DNA fragment size predicted by the McrBC (left panel) and HELP (right panel) enzyme digestions. (B) For all three methods, a 500-bp window was formed around each probe, the observed-to-expected ratio of CpG was calculated, and box-plots of the M-values are displayed by these ratios. Only probes related to fragments of sizes between 50 and 600 bp for McrBC, and between 600 and 1200 bp for HELP, are included.











