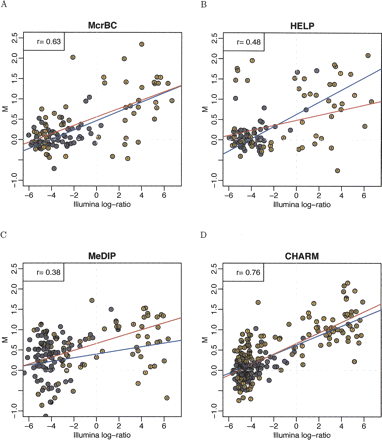
Comparison of method-specific methylation measurements to reference data. For the HCT116 (brown) and DKO (gray) samples, M-values from high-throughput methods are plotted against M-values from the Illumina reference platform. To illustrate the CpG observed-to-expected ratio, a 500-bp window was formed around each probe; this ratio (multiplied by 10) is displayed inside each point. A regression line was calculated and is displayed for probes with ratios <0.6 (blue line) and >0.6 (red line).











