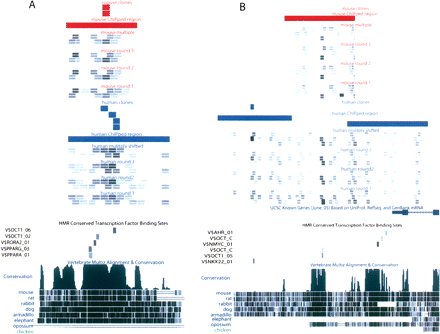
MEGAshift tracks for the UCSC Genome Browser. Two of the 19 genomic regions corresponding to REST (A) and GADD45G (B) have been displayed in custom UCSC genome browser tracks. Annotation is stacked vertically along the chromosomal coordinate axis (X-axis). Starting from the top and proceeding down, the mouse (red) sequences are annotated for the following molecules: oligonucleotides cloned out of POU5F1 selected fraction(short, stacked bars). ChIP-PET regions (wide bars), normalized enrichment scores in grayscale for each duplicate probe pair for each microarray experiment (multiply bound, round 3, round 2, round 1). Enriched oligonucleotides are shaded darkly. Human (blue) is identical save ChIPped material is analyzed by microarray (ChIP-chip). Predicted OCT binding sites annotated below. Conservation determined by eight vertebrate BLASTZ alignments.











