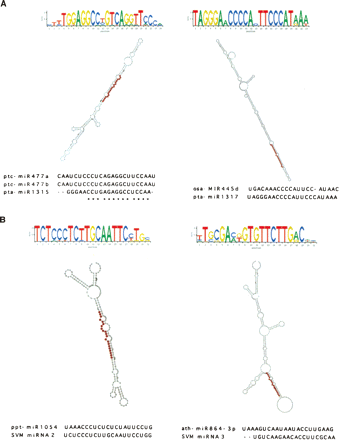
Summary of four novel miRNA clusters identified by two separate methods. Structures and sequence logos are shown for four novel miRNA clusters identified by either EST folding (A) or support vector machine (SVM) techniques (B). Sequence logos were generated from the multiple-sequence alignment of all small RNA sequences in the cluster. Folded structure (as predicted by RNAFold) includes positioning of the mature miRNA sequence (highlighted in red). The most abundant (highest count) sequence from each cluster was considered the most reliable representative mature miRNA and was used to search for EST (for those identified by SVM) and potential homologs in miRBase. The alignment of each of these sequences to either its best hit or hit(s) in miRBase is included.











