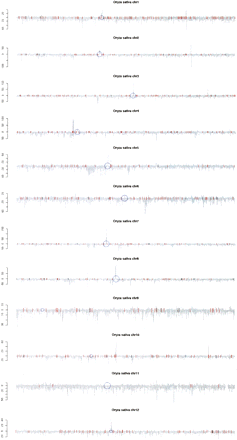
Degenerate alignment density of O. sativa 24- and 21-nt small RNAs to the nuclear genome. All distinct O. sativa small RNA sequences were aligned to all 12 chromosomes allowing degenerate alignments (see Methods). The 24-nt (positive axis) and 21-nt sequences (negative axis) demonstrated distinctive alignment patterns. Specifically, “hot spots” of 24-nt sequences include the heterochromatic regions including most centromeres (blue) as well as some clusters of ribosomal RNA genes (red). Positions of known miRNA genes are also marked (black).











