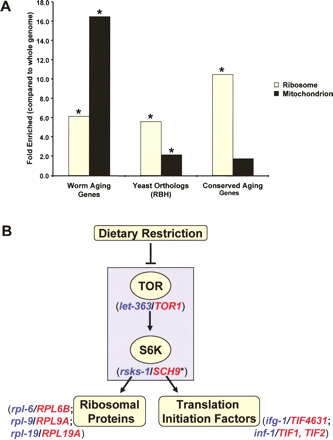
Conserved aging genes are enriched for ribosome-associated genes. (A) Gene ontology analysis of the 276 worm aging genes, their yeast putative yeast orthologs in the reciprocal best-hit set (RBH, gene total = 103), and the 11 conserved aging genes (Table 2) revealed a significant enrichment in the ribosome cellular component when compared with the entire worm or yeast genome ([*] P <0.05, after Bonferroni Correction for multiple comparisons). Mitochondrial genes are highly enriched in worm aging genes and their corresponding yeast homologs and orthologs, but there is no significant enrichment for mitochondrial genes in the group of conserved longevity genes identified in this study. Essential genes were included in the gene ontology analysis, but replicative life span was not measured for these (RBH/ribosome = 8; RBH/mitochondrion = 8). (B) Conserved aging genes highlight the role of the TOR signaling pathway in modulating longevity. Orthologous gene pairs are shown in parentheses (worm genes are in blue and yeast genes are in red). The nutrient sensing kinases are enclosed in the gray box. Many downstream targets of TOR signaling, including ribosomal proteins and translation initiation factors, play a conserved role in life span determination. (*SCH9 was recently shown to have S6K activity (Urban et al. 2007), but also has homology with worm aging genes in the Akt family of kinases, such as akt-1 and sgk-1.)











