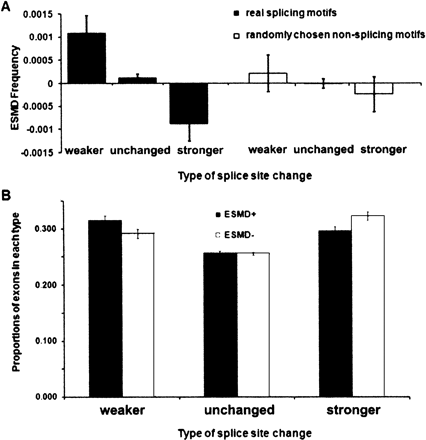
Compensation between splice-site changes and exonic splicing motif changes (comparing human to macaque). (A) Splicing-positive splice-site changes correlate with splicing-negative motif changes and vice versa. The exonic splicing motif difference (ESMD) is defined as the frequency of splicing-positive changes (ESE creations and ESS disruptions) minus the frequency of splicing-negative changes (ESS creations and ESE disruptions) in human relative to macaque. A positive ESMD is predicted to promote splicing while a negative ESMD would discourage splicing. Exons in the “weaker” set have one splice site at which the CV score (consensus values) has decreased by at least 5 on a CV scale of 0–100 and in which the other splice site has not increased by >5. Exons in the “stronger” set are defined in the opposite way. Exons in the “unchanged” set show no change at all in CV score for both 3′ and 5′ splice sites. The results of a control using non-ESE/ESS motifs, as described in the legend to Fig. 3D, are shown on the right. (B) Proportion of exons that have undergone changes in splice-site sequence and splicing motifs reflects compensation. Standard errors are indicated in both panels. The total number of exons in the weaker, unchanged, and stronger sets are 2897, 20,855, and 3394, respectively. Similar results comparing macaque exons to human are presented in Supplemental Fig. S3.











