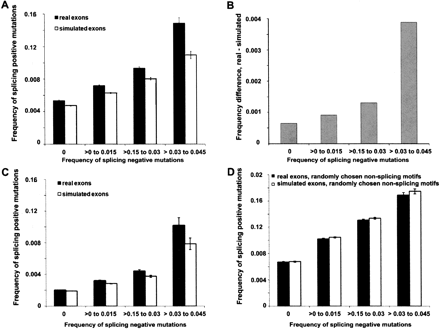
Evolutionary compensation of exonic splicing-negative motif changes by exonic splicing-positive motif changes. (A) Comparing human to macaque: exons were separated into four groups according to their frequency of exonic splicing-negative events (events per nt) as indicated. Exons with frequencies of 0–0.015, 0.015–0.030, and 0.030–0.045 generally contain 1 or 2, 3 or 4, and 5 or 6 splicing-negative differences, respectively. Exon pairs with frequencies higher than 0.045 have been omitted as many of them are poorly aligned. Black bars: real exons; white bars, simulated exons. (B) Net difference between the real and simulated sets is shown for each splicing-negative mutation category. (C) Same analysis as in A, except restricted to splicing changes in which the directionality of the change was determined by the use of dog as an out-group. Thus, in this panel the mutations in human exons can be considered ESE and ESS creations and disruptions rather than simply differences from macaque. (D) Same analysis as in A, but measuring differences using two randomly chosen non-overlapping sets of equal numbers of non-ESEs and of non-ESSs as controls. Similar results comparing macaque exons to human exons are presented in Supplemental Fig. S2.











