Data partition information
Click on table to view larger version.
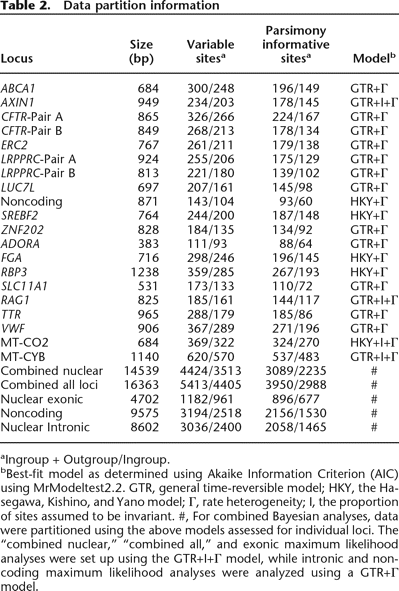
aIngroup + Outgroup/Ingroup.
bBest-fit model as determined using Akaike Information Criterion (AIC) using MrModeltest2.2. GTR, general time-reversible model; HKY, the Hasegawa, Kishino, and Yano model; Γ, rate heterogeneity; I, the proportion of sites assumed to be invariant. #, For combined Bayesian analyses, data were partitioned using the above models assessed for individual loci. The “combined nuclear,” “combined all,” and exonic maximum likelihood analyses were set up using the GTR+I+Γ model, while intronic and noncoding maximum likelihood analyses were analyzed using a GTR+Γ model.











