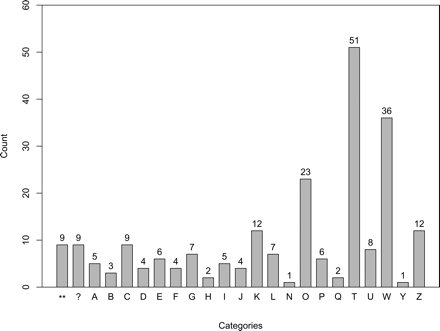
Distribution of promiscuous domains among functional categories of eukaryotic proteins. The categories are indicated with single-letter abbreviations on the X-axis, and the exact count of promiscuous domains in that category is shown on the top of each bar. If a domain is classified in more than one category, it is counted more than once. Abbreviations for functional categories (Tatusov et al. 2003) on the X-axis are: (A) RNA processing and modification; (B) chromatin structure and dynamics; (C) energy production and conversion; (D) cell cycle control, cell division, chromosome partitioning; (E) amino acid transport and metabolism; (F) nucleotide transport and metabolism; (G) carbohydrate transport and metabolism; (H) coenzyme transport and metabolism; (I) lipid transport and metabolism; (J) translation, ribosomal structure and biogenesis; (K) transcription; (L) replication, recombination, and repair; (N) cell motility; (O) post-translational modification, protein turnover, chaperones; (P) inorganic ion transport and metabolism; (Q) secondary metabolites biosynthesis, transport, and catabolism; (T) signal transduction; (U) intracellular trafficking, secretion, and vesicular transport; (W) extracellular structures and cell–cell signaling; (Y) nuclear structure; (Z) cytoskeleton; (**) various functions; (?) unknown function.











