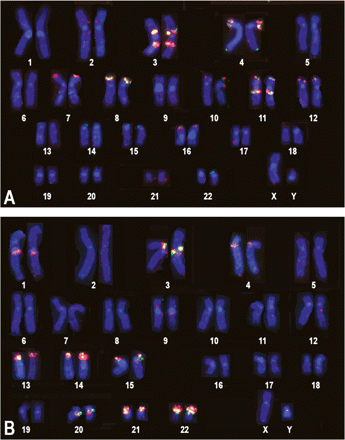
Two-color FISH of TBR1 probes on normal human chromosomes. DAPI staining (blue) based karyotype. (A) BACs RP11-71K3 and RP11-1053M22 (red in Supplemental Fig. 7E) give red and green FISH signals, respectively, at 3p12-p13, 3q21-q22, 4-pter, 7-pter, 7q21, 8-pter, 11-pter, 11q13, 12-pter, and 16-pter, corresponding to large TBSD. (B) BACs RP11-666K17 and RP11-139H7 (green in Supplemental Fig. 7E), give red and green signals, respectively, at 3p12-p13 and on the short arms of acrocentric chromosomes, suggesting that this region represents an “SD-amplicon” homologous to ribosomal-gene-cluster boundary (RBSD), which was not identified by sequence analysis. Both probes were hybridizing also to the pericentromeric regions of chromosomes 1, 4, 20, and 22, corresponding to SDs 6, 7, and 8 in Supplemental Fig. 7A–D.











