Table 2.
Summary of bacterial assemblies using 454 reads
Click on table to view larger version.
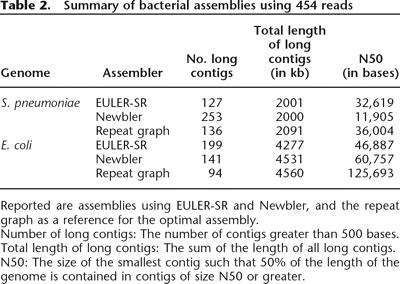
Reported are assemblies using EULER-SR and Newbler, and the repeat graph as a reference for the optimal assembly.
Number of long contigs: The number of contigs greater than 500 bases.
Total length of long contigs: The sum of the length of all long contigs.
N50: The size of the smallest contig such that 50% of the length of the genome is contained in contigs of size N50 or greater.











