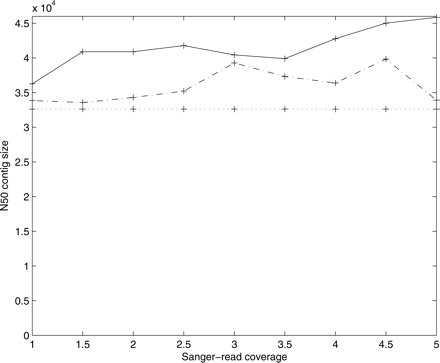
Figure 1.
Assembly of S. pneumoniae using 454 reads complemented with simulated Sanger reads (3000 ± 300 bp spacing). Uniformly distributed 800 base reads were generated for coverage varying from 1× to 5×, and then assembled using EULER-SR. We show the assembly using mate-pairs (solid), assembly using unpaired Sanger reads (broken), and the base assembly quality of 454 reads (dotted).











