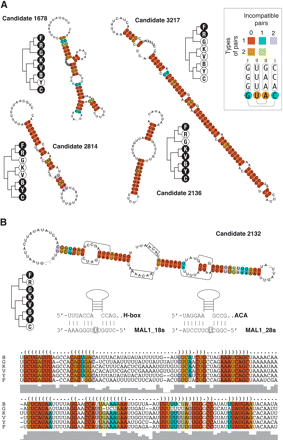
Examples of predicted structures for verified candidate RNAs. (A) Examples of structures of experimentally verified RNA candidates. Consensus structures are shown as described by Washietl et al. (2005a). Variable positions in stems are shown in circles. The number of different base pairs (in the alignment) supporting a given structure are indicated by colors (insert). An example is provided: Four columns from a hypothetical alignment involved in two base-pairings. Columns I contain one type of (correct) pairs (G:C), and one incompatible pair (G:A). Columns II contain two types of pairs, and no incompatible pairs. For each candidate, the phylogenetic distribution is indicated on a tree structure. Dark dots denote presence in a particular species (F indicates P. falciparum; R, P. reichenowi; G, P. gallinaceum; K, P. knowlesi; V, P. vivax; B, P. berghei; Y, P. yoelii; C, P. chabaudi). Candidate 3217 corresponds to the previously reported GC-rich repeats associated with var gene clusters (Hall et al. 2002). (B) Prediction 2132 is a H/ACA box snoRNA. H and ACA motifs are highlighted in boxes (the P. falciparum sequence has a degenerate H box) on the predicted structure; putative guide regions complementary to 18S and 28S rRNA are indicated by parentheses. Predicted complementarity to ribosomal RNAs is shown with target uridines (U673 and U3414 respectively) in boxes. The alignment of the six relevant Plasmodium species sequences is provided (b indicates berghei; g, gallinaceum; k, knowlesi; v, vivax; y, yoelii; f, falciparum).











