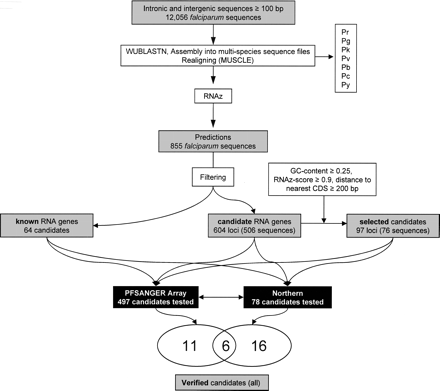
Flowchart of the prediction procedure, resulting in three classes of predictions; non-RNA predictions (non), known RNA predictions (known), and putative novel RNA predictions (candidates). Known RNA genes were identified from genome annotation and similarity to previously identified RNA genes (see in Methods). A set of candidates (selected) was chosen for Northern blotting analysis using the following criteria: (1) G+C content of the alignment >25%, (2) RNAz score ≥0.9, and (3) distance to nearest annotated protein coding gene >200 bases. Analyses and verification experiments are indicated by white boxes with black borders. Microarray indicates microarray analysis using the PFSANGER Affymetrix array; Northern, Northern blotting. (Pr indicates P. reichenowi; Pg, P. gallinaceum; Pk, P. knowlesi; Pv, P. vivax; Pb, P. berghei; Py, P. yoelii; Pc, P. chabaudi.)











