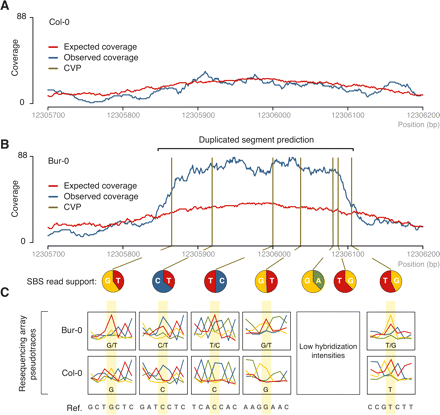
Detection of duplicated sequences using read coverage and sequence criteria. (A) Expected vs. observed read coverage in Col-0 for a region on chromosome 4 (positions are given at bottom). (B) Analogous region in Bur-0 harboring a predicted duplication. (Vertical lines) Seven CVPs in the region with elevated observed-to-expected coverage. The relative support for each base at a CVP is indicated below (pie charts). (C) Pseudotraces (see Supplemental Fig. S1 in Clark et al. [2007]) from resequencing array data, for five of the seven CVPs for which data was available (see Supplemental Methods). Compared with Col-0, double peaks are apparent in Bur-0 that match, in sequence, bases identified in the short-read data.











