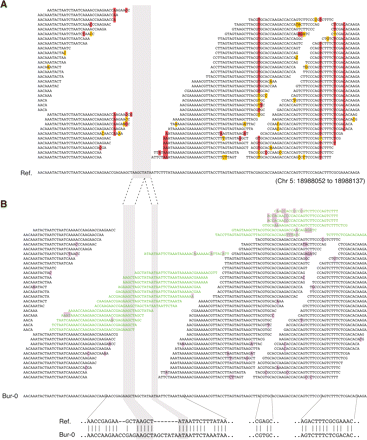
Targeted de novo assembly. (A) Example of alignment of Bur-0 reads to the reference (“Ref.”) sequence. Columns of high-quality mismatches (red) identify SNPs. A stretch of nucleotides without overlapping reads defined a target for de novo assembly (gray shading). Masked mismatches are highlighted in yellow. (B) Targeted-assembly derived Bur-0 contig for the same region, with reads added from the pool of unmapped (leftover) reads (green). Flanking SNPs identified in the mapping were recovered in the assembly, as was a complex sequence, which included two adjacent insertions and four SNPs in Bur-0 compared with the reference. The Bur-0 sequence was validated by PCR amplification and dideoxy sequencing. Mismatches to the contig sequence are highlighted in light purple.











