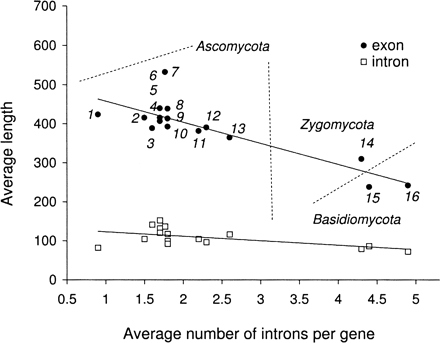
Annotations of the 16 fungal genomes demonstrate significant variability in gene organization: both in the average size of exons and introns as well as in the number of introns per gene (1, Schizosaccharomyces pombe; 2, Aspergillus niger; 3, Botrytis cinerea; 4, Stagonospora nodorum; 5, Fusarium oxysporum; 6, Magnaporthe grisea; 7, Neurospora crassa; 8, Fusarium graminearum; 9, Fusarium verticillioides; 10, Sclerotinia sclerotiorum; 11, Aspergillus terreus; 12, Aspergillus nidulans; 13, Coccidioides immitis; 14, Rhizopus oryzae; 15, Coprinus cinereus (also known as Coprinopsis cinerea); 16, Cryptococcus neoformans).











