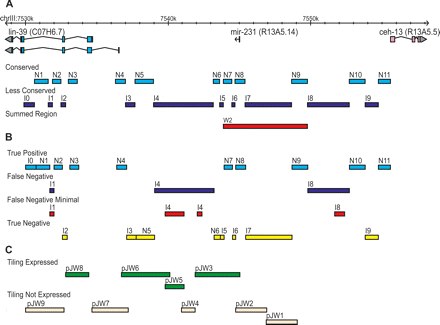
ceh-13/lin-39 Hox subcluster dissection based on sequence conservation. The ceh-13/lin-39 Hox locus was dissected into 21 sections for in vivo expression analysis based on the presence of MUSSA matches in a three-way alignment between C. elegans, C. briggsae, and C. brenneri. (A) MUSSA matches were used to identify similar, presumably conserved regions (N regions), which include the sequence match windows, 200 bp of 5′ and 3′ flanking sequence, and additional sequence for primer selection. The intervening, less-similar regions (I regions) located between the N regions were also tested. A “summed” region (W2) encompassing several component regions is shown as well. (B) With revised parameters of 100% match of 15-bp windows, the regions were repartitioned and true positives, true negatives, and false negatives were identified. The minimal region to recover the observed expression in the false negatives is identified (Streit et al. 2002; Wagmaister et al. 2006). (C) The regions assayed in the tiling analysis from Wagmaister et al. (2006) are shown for comparison, noting which drove expression (green) and which did not (beige).











