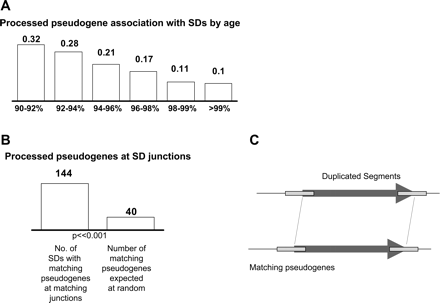
(A) Pseudogene association with SDs. Just like Alu elements, pseudogenes colocalize very strongly with old SDs and less so with younger SDs. All correlations are highly significant (P-value ≪ 0.00001). (B) Detailed SD junction analysis. A total of 144 SDs showed matching processed pseudogenes at both junctions, that is, both pseudogenes have the same parent gene and show high homology. When picking random genomic regions of the same size and number as SDs, no matching pseudogenes were ever found to overlap both SD junctions. When using a randomized offset of ±5 kb to account for potential sequence biases, an average of 40 matching pseudogenes were found, but in 1000 trials, never more than 43. (C) Schematic of matching processed pseudogenes at SD junctions. The processed pseudogenes overlap matching SD junctions at both duplicated segments, making them likely candidates for having mediated NAHR.











