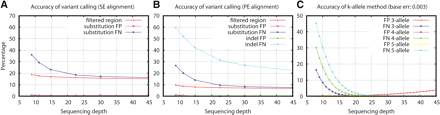
Accuracy of variant calling. In the figure, “filtered regions” are regions covered by three or fewer reads or by no reads with mapping quality higher than 60. For substitutions, FP equals the number of positions called as different from homozygous reference that in fact should be identical to the reference according to the simulation, divided by the total number of MAQ substitution calls; FN equals the number of positions that are different from the reference according to the simulation but are missed by MAQ, divided by the total number of mutations added in the simulation. For indels, FP equals the number of indel calls within 5-bp flanking regions of a true indel, divided by the total number of MAQ indel calls; FN equals the number of true indel calls missed by MAQ, divided by the total number of indels in simulation. (A) Variants are called based on single-end alignment. (B) Variants are called based on paired-end alignment. (C) Theoretical accuracy of k-allele method, where we call an allele as long as at least k reads are supporting the allele, assuming all reads are correctly aligned (see also Supplemental material).











