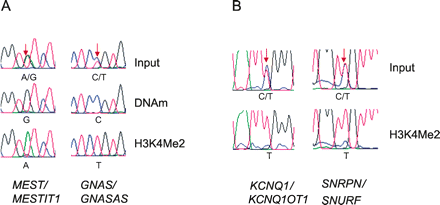
H3K4Me2 and DNAm double hits are allele specific. (A) Identification by SNP genotyping. The two marks are on separate alleles of the imprinted genes MEST and GNAS. H3K4Me2 ChIP and MeDIP were conducted on CEPH line GM10847, which was heterozygous for a SNP within the double-hit regions. The alleles are distinguished with polymorphisms, A/G (rs2301335) for MEST/MESTIT1 and C/T (rs6026560) for GNAS/GNASAS. Note that the A and G alleles are purified separately by ChIP and MeDIP for MEST/MESTIT1, and that the T and C alleles are purified separately by ChIP and MeDIP for GNAS/GNASAS. The paternal genotype is A/A for MEST/MESTIT1 and T/T for GNAS/GNASAS, consistent with paternal expression of the two genes. (B) Identification by serial analysis. Euchromatin was purified by H3K4Me2 ChIP, and the product was analyzed for DNA methylation by bisulfite sequencing. Methylation was assessed using a pseudopolymorphism at CpG sites, TG for unmethylated sites converted by bisulfite, and CG for methylated sites that resist conversion. For both the SNRPN/SNURF and KCNQ1/KCNQ1OT1 double hits, the input DNA is half methylated, while the euchromatin fraction is unmethylated. Arrows indicate polymorphic sites for A and CpG sites for B.











