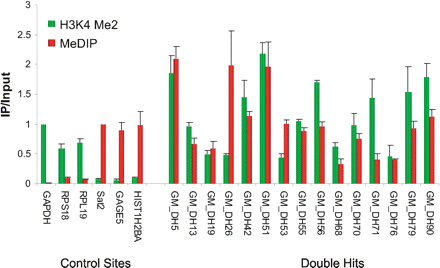
Validation of dual euchromatin/heterochromatin marks (“double hits”). Quantitative real-time PCR validation was performed for both chromatin immunoprecipitation (ChIP) with antibodies to histone H3 lysine-4 dimethylation (H3K4Me2), and methylated DNA immunoprecipitation (MeDIP) fractions, compared with input DNA. We analyzed three housekeeping genes, GAPDH, RPS18, and RPL19, as positive controls for H3K4Me2 (active), and two genes normally silenced in somatic cells, GAGE5 and HIST1H2BA, as well as satellite 2 repeats (Sat2), as positive controls for DNA methylation. The y-axis shows the ratio of ChIP or MeDIP to input. A value of 1 represents this ratio for ChIP of GAPDH and MeDIP of Sat2. All 15 double-hit sites discovered on the arrays were confirmed by this validation.











