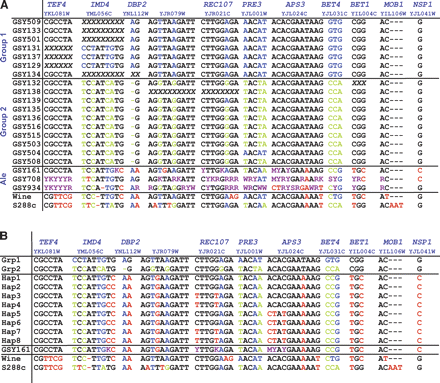
(A) Sequence differences between the Group 1 and Group 2 S. pastorianus strains (S. cerevisiae genomic portion only). Only nucleotides showing differences among the sequenced strains are shown. The five different gene regions sequenced are shown along the top; the primers used for both PCR and sequencing are given in Supplemental Table S1. Nucleotides in blue and green are those that distinguish Group 1 from Group 2 strains; nucleotides in red are those that are not shared with either Group 1 or Group 2 strains, and nucleotides in purple are those that are heterozygous in the ale strains, as determined from manual inspection of the sequence traces. Where a gene is not present in the genome because of deletion of that region of the S. cerevisiae genomic portion, the sequences are shown as a series of “X”s. Note that all of the heterozygous nucleotides or regions seen in the ale strains can yield (in at least one configuration) nucleotides consistent with the corresponding residues in the lager strains (either Group 1, Group 2, or both). (B) Sequences of the ale haplotypes inferred from spores of GYS161. As described, the ale strain GSY161 was sporulated and seven spores were sequenced across the 11 loci shown in the figure. For each unlinked locus with heterozygosities in the parent strain, unambiguous haplotypes were derived. Because there are three heterozygous loci, eight haplotypes can be inferred; all eight are shown in this figure with the one most similar to the Group 1 and Group 2 lager strains shown at the top of the list (“Hap1”).











