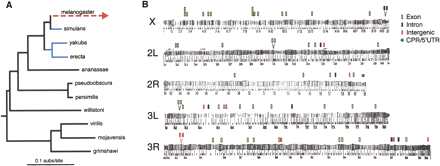
(A) Phylogeny of 11 Drosophila species with genome sequences. Branch lengths are derived from maximum likelihood analysis of all elements conserved throughout the tree. Branches in blue (D. simulans, D. yakuba, and D. erecta) were used to identify the blocks of at least 100 bp with 96% identity between the three species. All other lineages (and the D. melanogaster–D. simulans ancestor) were used to infer whether D. melanogaster had an accelerated rate of evolution relative to the expected rate of evolution based on elements conserved throughout the tree. (B) Locations of D. melanogaster accelerated regions (DMARs). (Stacked bars) Multiple DMARs within a single locus. (Two bars above a “V”) Two DMARs that were within the same chromosomal band. DMARs are found predominately in exons (46/64) and are significantly over-represented on the X chromosome (16/64). Chromosome images adapted from Lefevre (1976).











