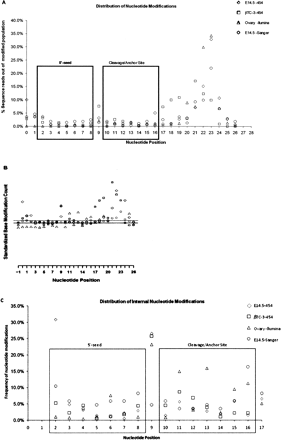
(A) Relative distribution of nucleotide modifications associated with mature miRNAs in relation to nucleotide position. This panel shows the distribution of nucleotide changes along the length of the mature miRNA. Modifications are highly suppressed in nucleotides 2–8 (containing the seed region that is important for miRNA–mRNA binding) and nucleotides 10–15 possibly defining additional determinants in the miRNA–mRNA binding. In sharp contrast, modifications are frequent at nucleotide 9 and the 3′ regions (nucleotides 16 and above). From these analyses we see that nucleotide positions 3–7 in the 5′-seed and nucleotides 10–15 exhibit significant suppression from nucleotide modification. (B) Statistical test for nucleotide modification distribution. This panel shows results from a statistical test used to examine the nature of nucleotide distribution in relation to nucleotide position of the mature miRNA. The method used is described in Methods. Under the null hypothesis where all positions behave the same with respect to base modification, we expect a random distribution of nucleotide modifications in relation to position. In this model we expect this quantity to be approximately normally distributed with a mean of 0 and standard deviation of 1. To ensure a simple plot, the median Z-score across all positions was set to 0 for each sample. The two black horizontal lines delineate the standard deviation around the Z-score. All data points above the upper line reflect modifications that occur at a rate greater than expected by chance. All data points below the lower line reflect modifications that occur at a rate less than expected by chance. From the data presented in this figure we see that the 5′-seed nucleotides 3–7, cleavage site nucleotide 10, and anchor nucleotides 14, 15 are constrained. Nucleotide position 9 is the most enriched internal position for modifications. (C) Relative distribution of internal nucleotide modifications. This panel shows the distribution of nucleotide changes in relation to internal positions (nucleotides 2–17). Modifications are highly suppressed in nucleotides 2–8 (containing the seed region that is important for miRNA–mRNA binding) and nucleotides 10–15 possibly defining additional determinants in the miRNA–mRNA binding. In sharp contrast, modifications are frequent at nucleotide 9 and the 3′-regions (nucleotides 16 and above). From these analyses we see that nucleotide positions 3–7 in the 5′-seed and nucleotides 10–15 exhibit significant suppression from nucleotide modification.











