MAKER’s performance on the C. elegans genome
Click on table to view larger version.
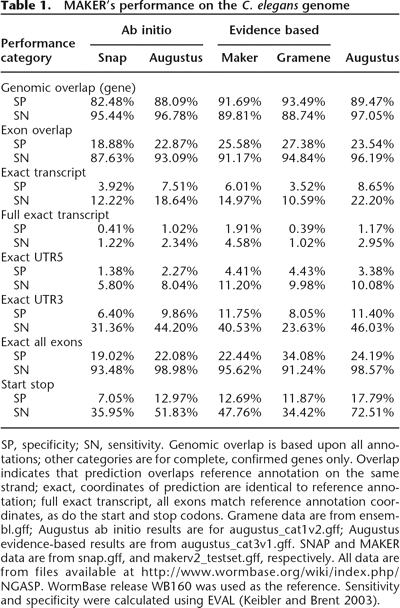
SP, specificity; SN, sensitivity. Genomic overlap is based upon all annotations; other categories are for complete, confirmed genes only. Overlap indicates that prediction overlaps reference annotation on the same strand; exact, coordinates of prediction are identical to reference annotation; full exact transcript, all exons match reference annotation coordinates, as do the start and stop codons. Gramene data are from ensembl.gff; Augustus ab initio results are for augustus_cat1v2.gff; Augustus evidence-based results are from augustus_cat3v1.gff. SNAP and MAKER data are from snap.gff, and makerv2_testset.gff, respectively. All data are from files available at http://www.wormbase.org/wiki/index.php/NGASP. WormBase release WB160 was used as the reference. Sensitivity and specificity were calculated using EVAL (Keibler and Brent 2003).











