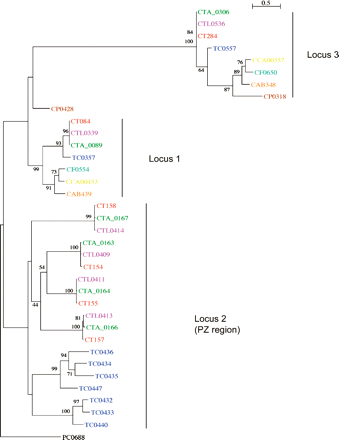
Phylogenetic relationships of chlamydial phospholipase D proteins. The protein names have been colored to indicate chlamydial strains: C. trachomatis strains: L2 (pink), UW-3 (red), and Har-13 (green); C. muridarum (dark blue); Cp. felis (light blue); Cp. caviae (yellow); Cp. abortus (orange); Cp. pneumoniae (brown); Candidatus Protochlamydia amoebophila UWE25 (black). Maximum likelihood tree built from protein sequences, using ClustalX, Phylip (Version 3.6), and NJplot. The numbers at the tree branches are percentage bootstrap values indicating the confident levels at that node where congruent. The bar indicates the genetic distance between species (one substitution per 100) as displayed in the branch lengths.











