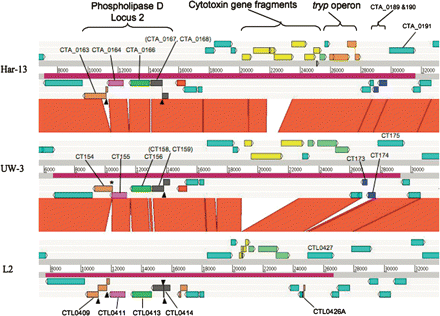
Comparison of the PZ locus of C. trachomatis strains L2, UW-3, and Har-13 ACT comparison (http://www.sanger.ac.uk/Software/ACT) of amino acid matches between the complete six-frame translations (computed using TBLASTX) of representatives of the PZ regions of the three sequenced C. trachomatis genomes: Har-13, C. trachomatis strain Har-13; UW-3, C. trachomatis strain UW-3; and L2, C. trachomatis strain L2. The red bars spanning between the genomes represent individual TBLASTX matches. Forward and reverse strands of DNA are shown for each genome (dark-gray lines). CDS are marked as colored boxes positioned on the three forward and three reverse translation reading frames (pale-gray lines). Regions mentioned in the text are marked and the CDSs labeled and color coded: Cytotoxin gene fragments (yellow), tryptophan biosynthetic genes (intact pale green; pseudogene orange), and phospholipase D (Locus 2; orthologous PLD CDSs are colored similarly). PZ regions are marked (purple box on DNA lines). The position of deletion events (*) and frameshift mutations (black arrowheads) within the PLD CDS are marked. Multiple systematic gene identifiers contained within parentheses indicates that the CDS was originally annotated as two CDSs.











