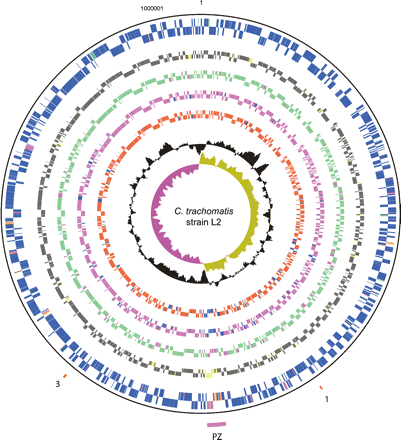
Circular representation of the C. trachomatis strain L2 chromosome. The outer scale shows the size in base pairs. From the outside in, circles 1 and 2 show the position of CDSs transcribed in a clockwise and anticlockwise direction. Using the published gene predictions for C. trachomatis strains UW-3 and Har-13, the strain L2 CDSs have been color-coded depending on whether they are: (blue) predicted and intact in all isolates; (pink) predicted and intact in L2 and UW-3; (green) predicted and intact in L2 and Har-13; (orange) defunct in L2, predicted and intact in Har-13 and UW-3; (red) unique to L2; (brown) defunct in all isolates. (Circles 3–10) C. trachomatis strain L2 CDSs that are present/absent in: C. muridarum (circles 3 and 4; present gray; absent yellow), Cp. felis (circles 5 and 6; present green; absent pink), Cp. caviae (circles 7 and 8; present pink; absent blue), and Cp. pneumoniae (circles 9 and 10; present red; absent blue) by reciprocal FASTA analysis. Circle 11 shows a plot of G+C content (in a 0.5-kb window); circle 12 shows a plot of GC skew ([G-C]/[G+C]; in a 0.5-kb window). The position of the PZ (pink) and the PLD CDSs in locus 1 and 3 are numbered accordingly and marked (red). Since the gene content of strain L2 and UCH-1 are essentially identical, these data apply equally to both isolates.











