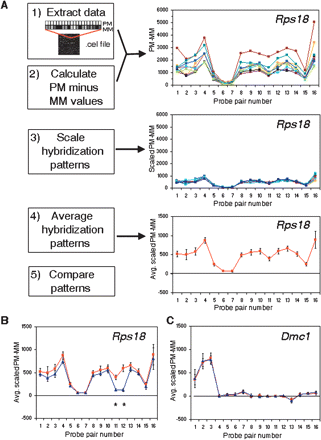
Detection of sequence differences using oligonucleotide array expression data. (A) Key steps in the GeSNP algorithm are described (left panels in boxes), and corresponding graphical illustrations of the SAMP10 data for MG-U74Av2 array probe set 98333_at, representing the gene ribosomal protein S18 (Rps18), are shown (right). Step 1 of the method is to extract data for a specific probe set from the CEL file. In step 2, the hybridization intensity difference between the perfect match and mismatch probe (PM − MM) for each probe pair (PP) is calculated. These values are then evaluated for inclusion in subsequent analyses as determined by passing pattern quality measures for detectable expression. The unscaled hybridization intensity values for Rps18 are shown for all nine samples of the SAMP10 strain, where the PP number is indicated on the X-axis ranging from 1 to 16, and the PM − MM value is shown on the Y-axis. Next (step 3), the intensity patterns for each sample are individually scaled to a common value. The scaled PP differences are then averaged (step 4) to generate a single value and standard deviation for each PP. (B) For the Rps18 probe set, the same analysis was performed for the nine SAMR1 samples, all of which passed the pattern quality measures for detectable expression. The average hybridization patterns with standard deviations obtained for SAMP10 (red line and squares) and SAMR1 (blue line and triangles) mice are shown. Using a t-value threshold of 6, the algorithm identified two PPs harboring putative sequence differences (black asterisks). Consistent with the hybridization pattern differences, DNA sequencing showed that each of these PPs indeed covered a region with a single base pair difference between the two strains. (C) The average hybridization signals with standard deviations are shown for the 96498_at probe set for the gene Dmc1, using the six files that passed the pattern quality measures for SAMP10 (red line and squares) and five files for SAMR1 (blue line and triangles). DNA sequencing identified no sequence differences between strains, consistent with the nearly identical, overlapping hybridization patterns (largest t-value of 2.4).











