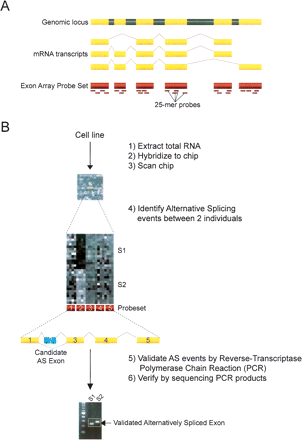
(A) Schematic for coverage of probe sets across the entire length of the transcript. Yellow regions are exons, whereas gray regions represent introns. The short dashes underneath the exon regions indicate individual probes of 25 nucleotides in length representing the probe set. The Affymetrix GeneChip Human Exon 1.0 ST Array allows for exon-level expression profiling in a single chip, and can interrogate over one million predicted exons within the human genome. (B) Flowchart for processing and analysis of chips to validation of alternative splicing events. Total RNA is extracted from the two cell lines (n = 15 replicates per individual) and is transcribed to cDNA and labeled with biotin. The total cDNA is then hybridized to the exon chip, followed by washing and staining with an anti-streptavidin antibody. Chips are then scanned, and hybridization data are processed and analyzed by the Affymetrix Power Tools (version 1.6) software package. A splicing index is calculated for ∼1.4 million probe sets covering one million exons. A subset of 20 alternative splicing events predicted between the two individuals using an unpaired t-test (P < 8.915 × 10−4) on the splicing index and other criteria (see Methods), are then validated by (1) RT-PCR using exon body primers flanking the probe set of interest and (2) sequencing of the RT-PCR products.











