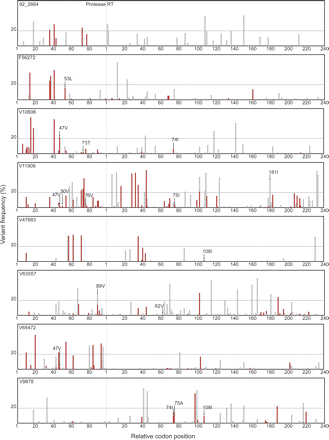
Sequence variants detected by ultra-deep pyrosequencing of eight clinical plasma samples including seven samples from antiretroviral-experienced patients (F56272, V10606, V11909, V47683, V63557, V65472, V9878) and one sample from an untreated patient (92_2664). Sequence variants were defined as differences from the consensus population-based sequence. The X-axis represents all 99 protease codon positions followed by the first 240 reverse transcriptase codon positions. Positions are demarcated at intervals of 20 codons. The Y-axis shows the frequency of each minor variant observed by ultra-deep pyrosequencing. Synonymous minor variants are shown in gray; non-synonymous variants are shown in red. The presence of more than one variant at the same position is indicated by superimposing variants of lower frequency onto variants with higher frequency. Drug-resistance mutations detected only by ultra-deep pyrosequencing are indicated with arrows. A horizontal line separates minor variants present at above and below a frequency of 20%. As noted in the text, 55 of 72 variants present in >20% of GS20 reads, but only nine of 392 variants present in <20% of GS20, were detected by conventional Sanger sequencing.











