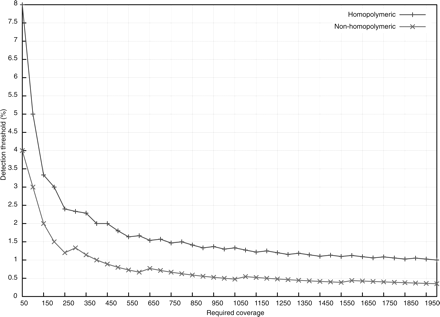
Relationship between the number of required ultra-deep pyrosequencing reads per position (Required coverage) and the detection limit of the frequency of a minority sequence variant (Detection threshold). The excess number of reads required to detect minor variants is a result of the Poisson model for handling potential sequencing errors. For example, ∼400 (rather than 100) reads are required to detect a minor variant present at about a 1% level in a non-homopolymeric region. Because homopolymeric regions are more likely to contain sequencing errors, higher numbers of reads are required to obtain similar sensitivities in these regions.











