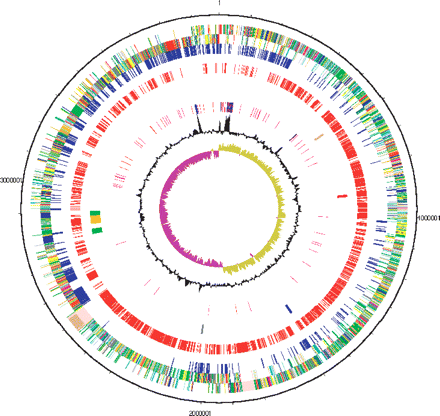
Circular representations of the genome of C. botulinum. The circles represent (from the outside in): (1 and 2) All CDS (transcribed clockwise and anticlockwise); (dark blue) pathogenicity/adaptation; (black) energy metabolism; (red) information transfer; (dark green) surface-associated; (cyan) degradation of large molecules; (magenta) degradation of small molecules; (yellow) central/intermediary metabolism; (pale green) unknown; (pale blue) regulators; (orange) conserved hypothetical; (brown) pseudogenes; (pink) phage and IS elements; (gray) miscellaneous. (3) (Blue) CDSs shared with sequenced clostridia. (4) (Red) C. botulinum-unique CDSs relative to the sequenced clostridia. (5) Virulence factors discussed in the text; (brown) streptolysin; (red) neurotoxins; (blue) six metalloproteases; (black) type IV pilus; (green) flagellar and chemotaxis operons; (orange) flagellar glycosylation island. (6) RNA genes; (blue) rRNAs; (red) tRNAs; (purple) stable RNAs. (7) G+C content (plotted using a 10-kb window). (8) GC deviation [(G − C)/(G + C) plotted using a 10-kb window]; (khaki) values >1; (purple) values <1.











