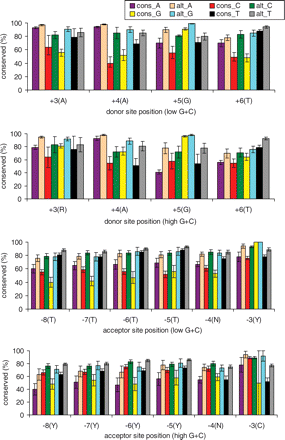
Mouse–human sequence conservation by splice-site position. cons_N: nucleotide N in human constitutive sites; alt_N: nucleotide N in human alternative sites, N = A, C, G, and T. Error bars represent 95% confidence interval. Consensus nucleotides are indicated in parentheses after each position. G + C content is calculated from the 10-kb region around human splice sites; 44% is the threshold defining low and high. Only paired, “nonexonic” sites (those not found in the interior of an exon in any spliced variant) were used.











