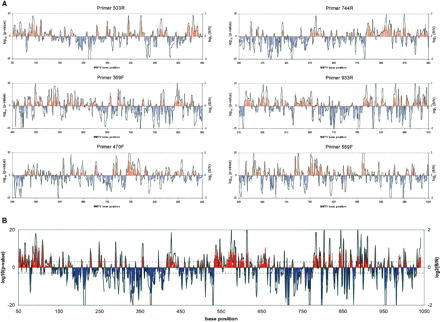
Primer extension data for the MMTV-LTR from each primer and their combined result. (A) The PCR extensions from each primer were analyzed to determine locations of significant difference between the bare genomic and nucleosomal DNA (B and N, respectively). The 5′-start position of each primer is given as well as the strand it primes from (F, forward; R, reverse). In each graph, the red and blue lines indicate the locations and log10(P-value) of significant differences between the two, while the black line shows the log2(intensity ratio) of the mean intensities for each trace. The dotted lines represent a P-value of 0.001, and have been included solely as a reference point for the reader. Above midline indicates B > N, below midline indicates N > B. (B) Combined information from all six primers to reveal MNase accessibility across the MMTV-LTR.











