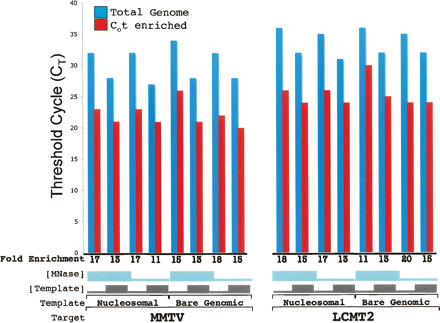
Quantitative PCR of total genomic and Cot-enriched DNA samples. Quantitative PCR was used to confirm enrichment of MMTV and LCMT2 target DNA. Nuclei from MDA-kb2 cells (nucleosomal) and chromosomal (bare genomic) DNA were digested with two concentrations of micrococcal nuclease as described in Methods. DNAs were isolated, melted, and allowed to rehybridize to a Cot value of ∼3332.2 M·sec. The slowly reassociating single-stranded component was purified by hydroxyapatite chromatography as described in Methods. Five or 50 ng of this Cot-enriched DNA (red columns) was compared with 5 or 50 ng of the non-Cot-enriched starting material (blue columns) in a quantitative PCR experiment assessing the enrichment of two loci, MMTV and LCMT2. The fold enrichment between the total genomic sample and the Cot-enriched sample was calculated using an amplification efficiency of 1.84. The value shown is CT, the fractional cycle number at which the amount of amplified copies reaches a fixed threshold.











