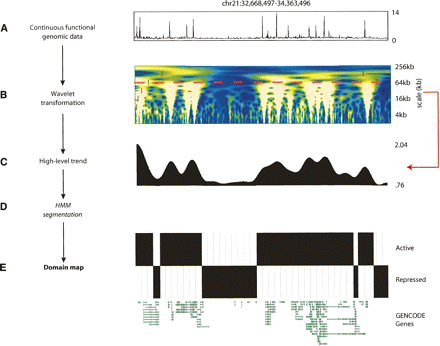
Wavelet segmentation approach for functional domain mapping. (A) Exemplary continuous functional data type (H3 acetylation) for ENCODE region ENm005. (B) Continuous wavelet transform heatmap (“scalogram”) of H3 acetylation data. In the heatmap, the horizontal axis represents genomic position, while the vertical axis represents wavelet scale. Each color in the scalogram represents the magnitude of the wavelet coefficient at that genomic position and scale, ranging from blue (small magnitude) to white (large magnitude). Larger magnitude wavelet coefficients imply a strong trend in the original data at that position and scale. The 64-kb scale is marked with a dashed red line. (C) Wavelet smoothed data at the 64-kb scale obtained using MODWT. Horizontal axis: genomic position. Vertical axis: wavelet coefficient at that position at the 64-kb scale. (D) Results from two HMM state segmentation of data from C, based on fitting HMM to H3ac data over all ENCODE regions. The top row indicates state 1 regions in black, while the bottom row indicates state 0 regions in black. The high state (state 1) is taken to represent active domains based on the assumption that H3ac is an activating mark. (E) GENCODE gene annotations for ENm005. Note the correspondence between state 1/active and GENCODE gene and density.











