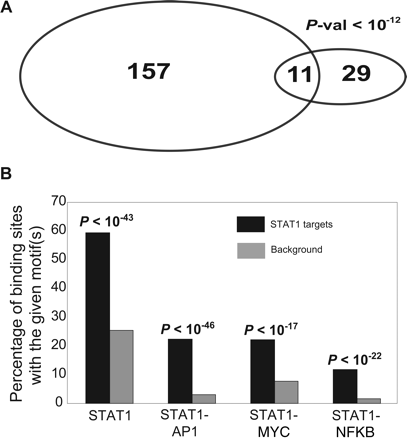
(A) Overlap of STAT1 target genes identified by STAGE with STAT1 target genes identified by ChIP-chip using a core promoter array. STAGE identified 29 promoters out of the ∼9000 promoters present on the core promoter array as STAT1 target promoters. Eleven out of these 29 overlapped with the 157 promoters identified as STAT1 targets by ChIP-chip analysis at an enrichment ratio greater than threefold. The enrichment ratio refers to the ratio of the fluorescence intensity of ChIP DNA to that of reference DNA at each spot on the core promoter microarray. (B) Motif analysis. The Y-axis shows the percentage counts of the number of sites bearing the given motif(s) out of the 381 STAT1 binding sites detected by STAGE. Almost 60% of the 381 binding sites had the STAT1 motif TTCNNNGAA as compared to 27% in the background. We also detected an enrichment for the co-occurrence of the binding motifs for STAT1 and AP1 (TGAG/CTCA), STAT1 and MYC (CACA/GTG), and STAT1 and NFKB (GGGA/GNNC/TC/TCC) in accordance with the fact that STAT1 exhibits cooperative binding with these factors to regulate downstream promoters.











