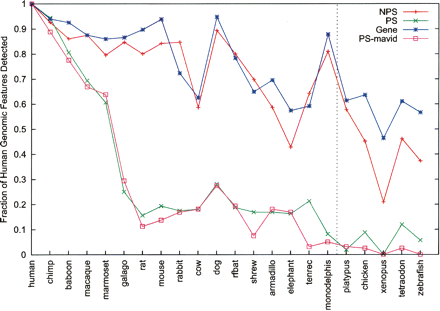
Preservation of human genomic components in other species. The number of human pseudogenes (or genes) with orthologous sequences in individual species was computed and then plotted (by normalization with the total number in human) against each species. Only exons (or pseudoexons) were used in these analyses; (NPS) nonprocessed and (PS) processed pseudogenes. Data were derived from sequence alignment constructed by the program TBA except PS-mavid, which was by MAVID. Note that species with sequences available for the ENm001 region only are omitted in this figure. A more comprehensive plot (of this figure and also Fig. 5A) with data for introns and other genomic data can be found in Supplemental Figures S1 and S2. The data for non-mammalian species (right of the vertical line) should be taken with more caution because ortholog assignments for these species are likely more difficult.











