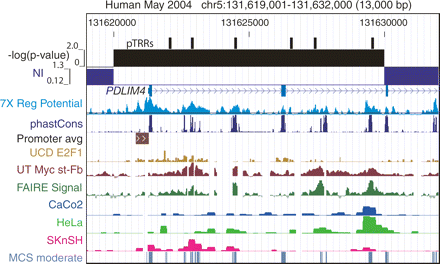
Recent positive selection in the PDLIM4 gene. The customized view from the UCSC Genome Browser covers about 13 kb of ENCODE region ENm002 centered at gene PDLIM4. It shows the locations of pTRRs, the P-value for deviation from neutrality for 10-kb windows (H. Lawson, J. Martin, D.C. King, B. Giardine, W. Miller, and R.C. Hardison, in prep.), the neutrality index (Rand and Kann 1996; with values >1 implying negative selection and values <1 implying positive selection), positions of known genes, regulatory potential scores (Taylor et al. 2006), phastCons scores (Siepel et al. 2005), active promoters (Cooper et al. 2006), occupancy by E2F1 (Bieda et al. 2006), occupancy by c-Myc (Kim et al. 2005), formaldehyde-assisted isolation of regulatory elements (FAIRE) (Giresi et al. 2007), DNaseI hypersensitive sites measured in CaCo2, HeLa, SKnSH cell lines/phenotypes (Sabo et al. 2006), and moderate MCSs generated by the ENCODE Multispecies Analysis group (Margulies et al. 2007).











