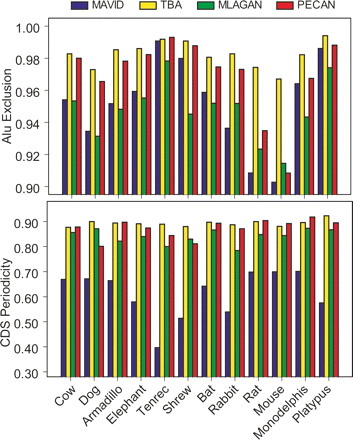
Alignment “correctness” as measured by Alu exclusion and periodicity of substitutions in coding exons. For a group of non-primate mammals, the fraction of human Alu bases (out of a total of ∼3.8 million) that are not aligned (i.e., gapped) is shown (top panel). A score of 1 would correlate with complete exclusion of all Alus, as would be the case in alignments with no false-positive orthology predictions. We also show the fraction of human coding exons that show a triplet periodicity in substitutions in the pairwise alignment between human and each query species (see Methods). Note that this is purely a relative measure, since we exclude exons that are completely gapped in at least one alignment, or fail to show periodicity in at least one alignment.











