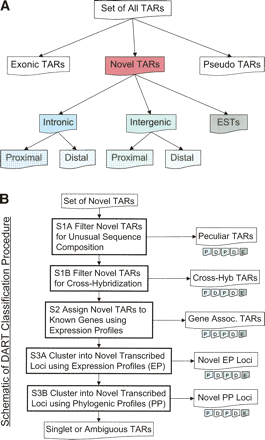
(A) Schema for the partitioning of TARs on the basis of location relative to GENCODE genes and pseudogenes (also see Table 1). Proximal regions are located within 5 kb of the nearest GENCODE exon. (B) Outline of the DART classification procedure of novel TARs. Novel TARs are first filtered on the basis of sequence composition (step 1), and then a fraction of the remaining novel TARs are associated with known genes (step 2). A portion of the remaining novel TARs are clustered in novel transcribed loci on the basis of expression profiles (EPs) and phylogenetic profiles (PPs) (step 3). See Table 2 for the numbers of novel TARs classified by each of these steps. The singlet and ambiguous TARs are what remains at the end of the classification procedure.











