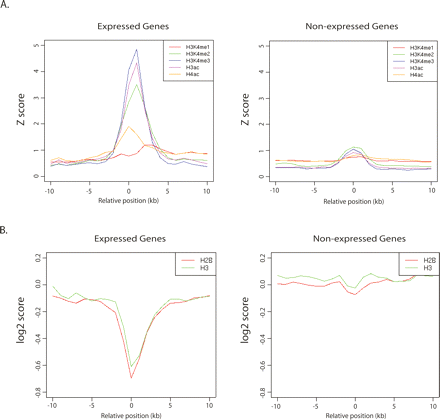
The histone modification profile at TSSs for active and inactive genes. (A) The average Z-scored histone modification profile for ±10 kb surrounding each TSS of a gene split according to the expression level of the gene as determined by the GeneSpring MAS5 present (Expressed) and absent (Non-expressed) analysis of Affymetrix U133 plus 2.0 gene expression data from the GM06990 cell line. Histone modification signals are plotted as lines for the average Z-score over all HMM sites in each class with lines color-coded as in Figure 4A. (B) For comparison this plot shows the mean log2 value of the enrichment for ChIP-chip using antibodies to histone H3 and H2B over all ENCODE TSSs, split according to the expression level of the gene as determined by the GeneSpring MAS5 present (Expressed) and absent (Non-expressed) analysis of Affymetrix U133 plus 2.0 gene expression data for the K562 cell line.











