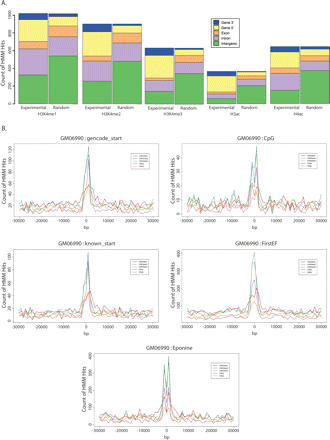
Distribution of histone modification sites in the lymphoblastoid cell line, GM06990. (A) The plot shows the number of histone modification sites identified by the HMM which overlap with an exon (orange), intron (mauve), gene 5′ end (yellow), gene 3′ exon (blue), or intergenic sequence (green) for the lymphoblastoid cell line, GM06990, in the ChIP-chip data (left panel) or in random simulated data (right panel). Random data were simulated by generating sites of the same size distribution as the experimental data and placing them at random among the ENCODE regions represented by the PCR product microarray. This was repeated 100 times, and the mean frequencies of overlap were plotted. (B) Distribution of histone modification sites with respect to gene starts. The distance to the nearest gene start for histone modification sites identified by the HMM relative to GENCODE annotation (gencode_start), UCSC known gene starts (known_start), CpG islands (CpG), FirstEF gene start predictions (FirstEF), and Eponine predictions (Eponine) was determined. The plot shows the frequency of distances to the nearest gene start for H3K4me1 (red), H3K4me2 (blue), H3K4me3 (green), H3ac (mauve), and H4ac (orange) in 1-kb windows over ±30 kb from a start.











