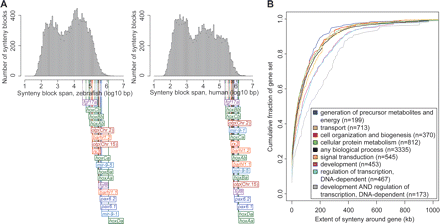
The studied loci are within large synteny blocks. (A) Histograms show the span of synteny blocks in the zebrafish (left) and human (right) genomes. Colored lines indicate the genomic spans of synteny blocks for the loci investigated in this study and, for comparison, the loci of the seven zebrafish hox clusters. Note that zebrafish gene symbols are used in both histograms in order to differentiate between synteny blocks that overlap on the human genome (e.g., the pax6.1 and pax6.2 synteny blocks, which partially overlap at the human PAX6 locus). Synteny blocks were computed based on alignments between the two genomes as described in Methods. (B) Each curve shows the cumulative distribution of extent of synteny around genes participating in a particular biological process. The distributions for the processes transcriptional regulation and development grow significantly slower compared to any of the other processes investigated (P < 0.004; one-tailed Kolmogorov-Smirnov test). For genes involved in both of these processes, the difference is highly significant (P < 1 × 10−6). Numbers within parentheses indicate the number of genes annotated to a process and located within a synteny block.











